test account: test Password: bio123456789
| chromosome length | 2A 780798557 3A 750843639 7B 750620385 4A 744588157 7A 736706236 6B 720988478 5B 713149757 5A 709773743 1B 689851870 4B 673617499 2D 651852609 7D 638686055 6A 618079260 3D 615552423 1A 594102056 5D 566080677 4D 509857067 1D 495453186 6D 473592718 |
| genomic location | chrom pos type 2A 50602155 A 2A 58861277 A 3A 178099541 A 3A 77794217 B 4D 20732565 B 4D 11935999 B |
| Output |
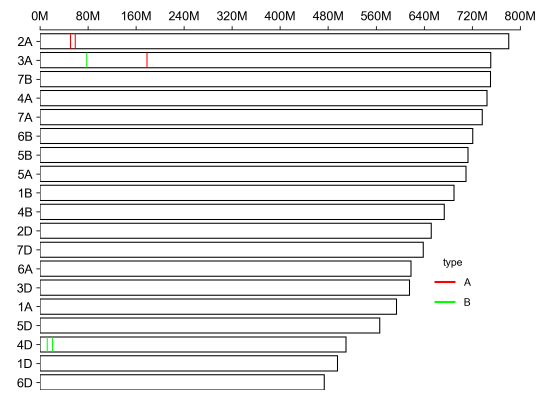
|
1) How to plot?
1, Put data in excel according to the example format.
2, Copy and paste into input frame.
3, Input pre-checking button to check input
4, After checking pass, select parameters, submit and download
2) How to cite?
3000+ papers in (Google Scholar)
Tang D, Chen M, Huang X, Zhang G, Zeng L, Zhang G, Wu S, Wang Y. SRplot: A free online platform for data visualization and graphing. PLoS One. 2023 Nov 9;18(11):e0294236. doi: 10.1371/journal.pone.0294236. PMID: 37943830.
3) FAQs